|
Week 20 consisted of completing experimentation and wrapping up the data collection. All data is viewable on the google sheet and it corresponds with the information and experiment details on the daily experimentation blog.
DATA SHEET: https://docs.google.com/spreadsheets/d/1Z5OjVgMMSBnSf9DHKzgpji-Gk7PT_VH5ukTOlYvM4Os/edit?usp=sharing DAILY EXPERIMENTATION BLOG: https://docs.google.com/document/d/1z4J1TUco5qNMQIhQyP3dqujcpUmezmx677Hhu3eyB30/edit?usp=sharing
0 Comments
Week 19 consisted of gathering data using the pond water as the growth media as opposed to the nutrient broth. This was done to see the difference between a so-called "perfect" scenario (the nutrient agar) and an imperfect "real" scenario (the pond water) it is a little difficult to see any results simply from the images as all the non-coliformal colonies grew up as well on the plate, however, we counted it against a control plate of un-infected pond water growth to count out any non-E.Coli B colonies.
For week 18, the dilution set assay was re-run to prove its accuracy and correctness. The secondary results were just as promising as the first, with significant killing of E.Coli B seen in all dilutions, with a cleared, or almost cleared, solution in the 50, 5, and .5 cells/mL dilutions which are the most equivalent of an actual lake environment (shown in pictures 4-9). In the coming weeks after spring break, we will be running these assays with the actual pond water to see if the results are re-creatable in the real-life solution in which the interaction would be taking place. Also, distribution and delivery methods will be theorized and possibly tested before wrapping up experimentation.
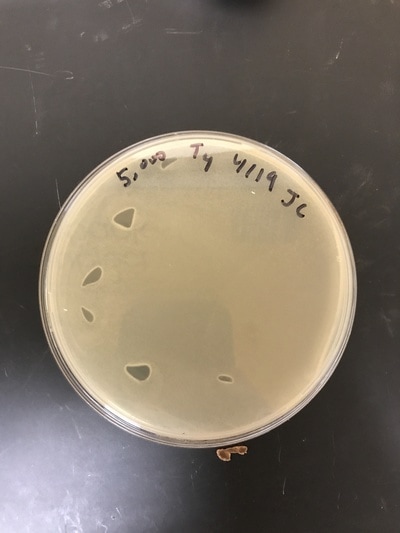
After the successful run of the serial dilution method for the E.Coli B colonies, the same serial dilution method was used (beginning with the start concentration of 50,000 cells per mL) and combined with the two phage cultures T2 (Pic 1) and T4(pic2). Similar results to the control serial dilution can be seen; however, when combined with the phages, the dilutions of 50, 5, and .5 cells per mL have noticeable less colonies, almost none. These are promising results. The experiment will be run again for confirmability and the addition of more time points to gather more data on how the phages kill. Data was entered into the spreadsheet for further data analysis. Also, time will be taken to gain a deeper understanding of Michaelis-Menten Kinetics to enrich the understanding of the project, our results, and conclusion.
For week 16, another dilution series was run to get baseline data for verify the validity of the dilution method being used to plate the E.Coli B colonies. Using the flamed bent glass method (pic 1) 12 plates were inoculated with a series of increasingly dilute E.Coli broth cultures using the serial dilution method where a 1:9 ratio is used of infected:clean broth in succession to reach the desired dilution range. To get a good starting point, a cytometer slide (pic 3) was used to count cells in the original culture and when only one cell of E.Coli could be seen the broth had a concentration of 50,000 cells per mL. Results analyzed after sufficient days of growth showed success of the dilution process, all colony counts were entered into the spreadsheet for later data analysis.
For week 15, it was decided that a new set of all strains needed to be plated. After re-plating the specimens, the newly-filtered stock of bacteriophages, both strain T4 and T2, were tested for contamination. The phages were also tested for viability; a small sample of the phage and E.Coli broth was taken out of one of the sterile filtration setups before it was filtered through, it was plated against some broth of pure E.Coli (picture 1) and as can be seen in the picture, no E.Coli grew, indicating a positive result. To get a set of baseline data for the core experimentation, a gradient of different ratios of phage:E.Coli were grown in a series of snap tubes and plated out to see how many colonies grow based on the ratio of phage to culture (photos 2-6). These results will be gathered and tweaked to get a solid set of baseline data for the core of the experimentation.
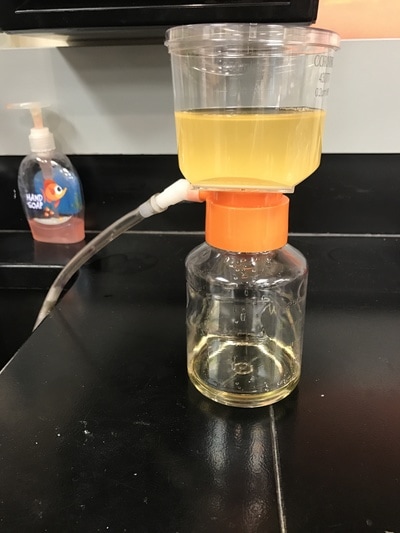
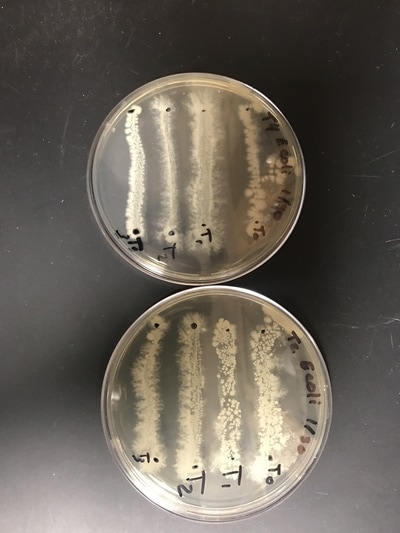
With week 14 we continue research after the earlier contamination setback. As can be confirmed from photos 1-3, the colonies re-grown after the contaminated colonies were disposed of are pure E.Coli. We have obtained and are now working with a new E.Coli strain as well: E.Coli strain B, with hopes that the new strain will be a more prolific host for the phages and produce better results faster and more effectively. A bubbler setup was created (Photos 4, 5) to grow the phages, again, in hopes that the process will be more productive and give stronger results and phage output. The last step of the process to purify phages was to run the phage/E.Coli broth through a sterile filter mechanism to filter out E.Coli cells and cell parts and get a pure phage stock (Photo 6).
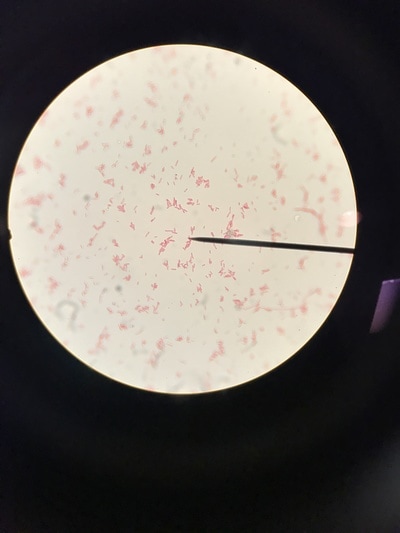
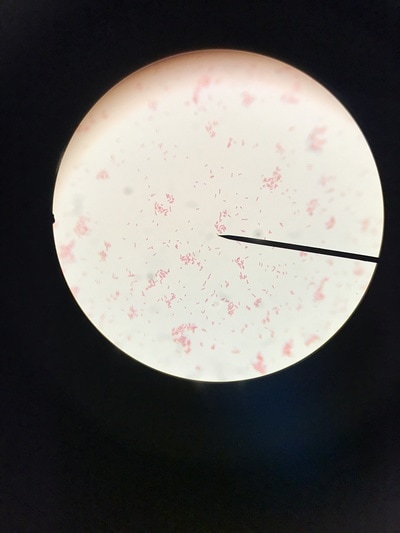
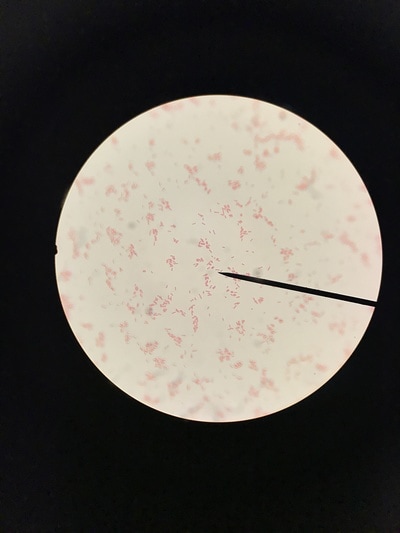
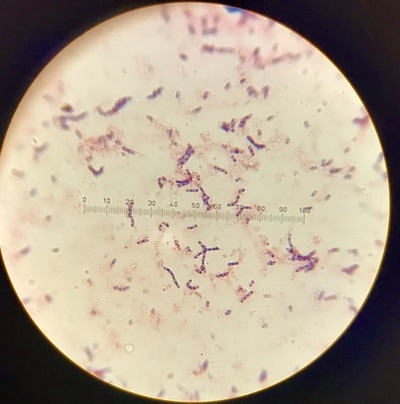
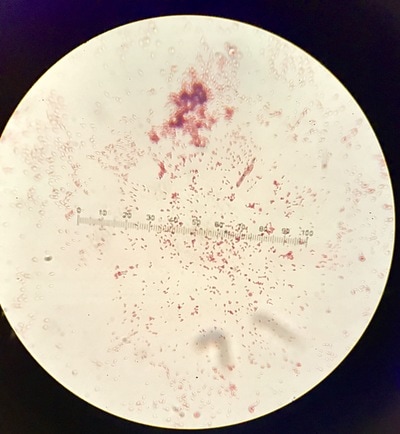
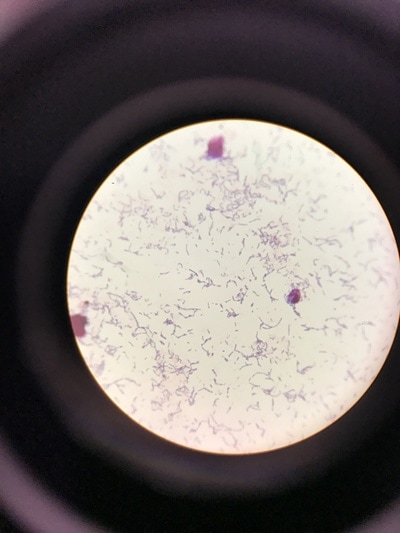
Week 13 is a continuation of growth of E.Coli and testing for killing using the phages. A small setback occurred during this round of testing that involved a contamination problem. It was found that the E.Coli stock we had been working with had gotten contaminated somewhere along the way and the reason we weren't seeing as much killing of our culture was because it was the wrong culture and the pages aren't built to kill it. After gram staining (pictures 1,2, and 5) we could see that instead of a gram-negative individual bacillus (pink, individual, rod-shaped cells) we were looking at a gram-positive strepto-bacillus (purple, chain-linked, rod-shaped cells). Unfortunately thus is the nature of microbiology and bacteriology; when working with subjects and organisms this small and numerous often contamination occurs and there is no clear source or fault. It just happens. Despite this, we will kill all contaminated sources and continue onward with experimentation. Setbacks happen, but all we can do is learn from this and press forward.Week 12 harbors the first bold steps into the heart of the experimentation; the investigation of the core hypothesis. The week began by creating large broth cultures of E.Coli to later be innoculated with the T2 and T4 phage cultures so that the phages may replicate within the E.Coli bacteria and we may retrieve a stock of phage solution for later use. The broths were created and set to grow in the incubator. Before the E.Coli broths were turbid enough to use to make the stock solutions, a small sample was taken and placed in two snap tubes, one was innoculated with T4 phage culture, and one with T2. To keep the solution moving and viable, a contraption was innovated which involves a stir bar and a small amount of water to agitate the snap tubes since we unfortunately didn't have a shake plate at our disposal. These will grow and be used as a preliminary test set to see how effective and efficient the phages are at ridding solutions of E.Coli contamination. The contents of the snap tubes were streaked out after many hours and the growth will be analyzed in the coming days. With the information gained from these experiments we will edit the rest of our procedure for the experimentation to fit our findings and streamline our testing. Week eleven has been a lot of preparation. While these steps are extremely important, it has been a lot of the same activities. We began by pouring agar into the amount of plates that will be carrying us through the first few rounds of experimentation (pictured below). This was accomplished by a day of mixing and cooking the agar recipe, autoclaving (sterilizing) it and pouring the actual plates out. Along with plates, we poured about 30 agar slants in which we will grow the initial E.Coli colonies. The tank continued to settle in and get re-oxigenated via bubbler (pictured), the established flora and mineral levels will continue to stay within normal pond levels as long as there is some form of oxygenation and current. We streaked the E.Coli onto the new slants, as we will be doing every 3-4 days to keep them fresh, and made sure the incubator was working; it was, and the E.Coli are prospering. Due to midterms, we had to work our schedule around the coming week so there will not be any active, or major, experimentation, and therefore no blog updates, until the week after next. You can stay up to date with a daily account of all laboratory activity by viewing the team's Daily Experimentation Log (Link on the homepage).
|
































































 RSS Feed
RSS Feed
